Elisha Krieg Group
DNA Nanotechnology: building programmable materials and nanometer-scale devices

Synthetic polymer materials have revolutionized the world within the last century. Yet, a glance at nature reveals, how limited the functions of synthetic polymers still are in comparison to biological matter: biological cells use molecular self-assembly and self-organization to build complex materials (e.g. the cytoskeleton) and to operate sophisticated macromolecular devices (e.g. motor proteins). For many years, chemists and material scientists have been intrigued by one question: How can we construct materials with equally sophisticated and versatile features?
Inspired by biology’s refined design principles, our group strives to develop materials with programmable and reconfigurable properties. We use DNA as the quintessential component for these materials: aside from its crucial role as the carrier of genetic information, DNA can be used as a construction material to assemble artificial nanometer-scale structures. This research field at the interface of science and engineering is known as DNA Nanotechnology.
By combining concepts of DNA Nanotechnology with traditional polymer chemistry and molecular biology, we aim to transfer the programmability and reconfigurability of nanometer-scale DNA devices to the macroscopic level. To this end, structural scaffolds of synthetic acryloyl polymers are endowed with DNA-based nanomodules via non-covalent self-assembly (see Figure). The ultimate target is the development of new classes of materials that can be programmed to perform complex tasks for diverse biomedical applications (see Future projects and goals).
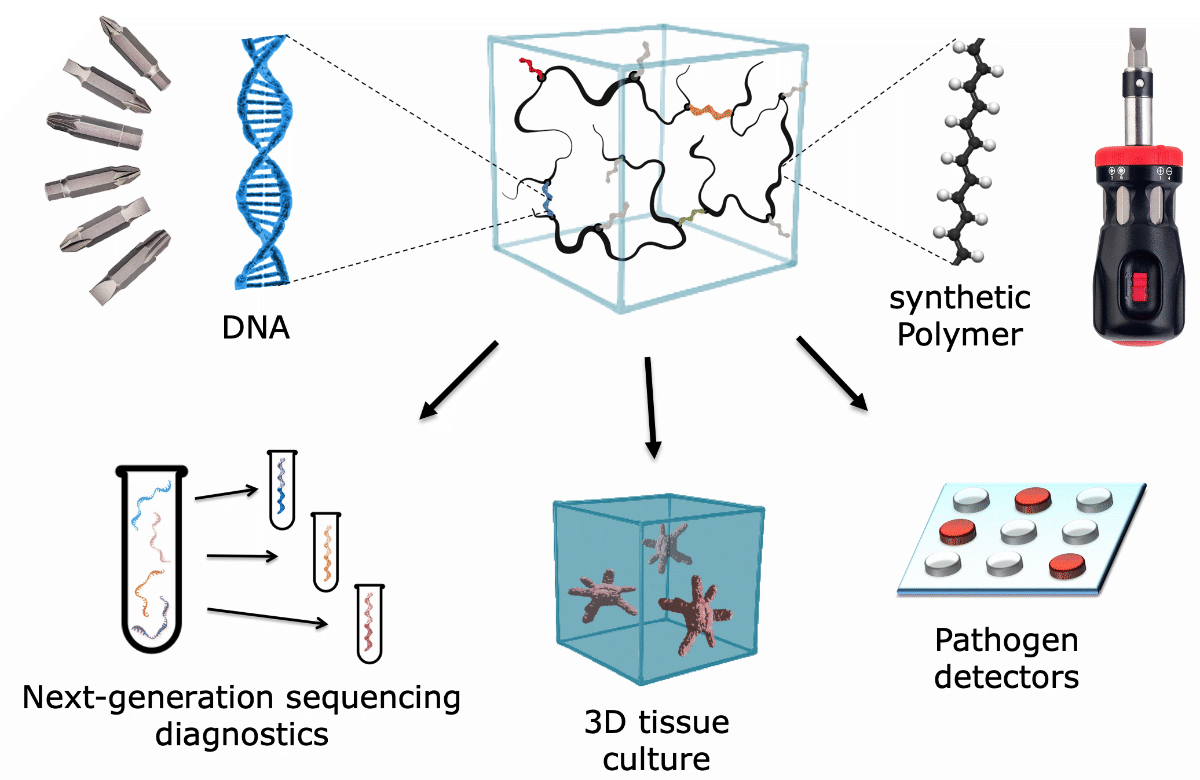
Future Projects and Goals
Our future research will focus on the development of three application areas, emerging from a single programmable material platform (see Figure):
- Materials for purification and labeling of DNA and RNA libraries will increase the efficiency and sensitivity of new diagnostic assays that rely on Next-Generation Sequencing.
- Printable and dynamically reconfigurable 3D cell culture matrices will be used to generate better in-vitro models of healthy and damaged tissues.
- The point-of-care detection of pathogen-specific nucleic acids will be achieved by materials that sensitively respond to DNA-based reaction networks.
Methodological and Technical Expertise
- DNA Nanotechnology (sequence design, assembly, DNA origami)
- Organic, supramolecular and polymer chemistry
- Standard methods in molecular biology
- Nanoparticle synthesis and purification
- Nanomaterial characterization: electron microscopy, atomic force microscopy, and oscillatory rheology