Rolf Jessberger Group
Chromosome Dynamics and Hematopoietic Cell Biology

© Stephan Wiegand
My lab focuses on three major areas:
- Cohesin and cohesin-associated factors and their contribution to genome integrity, chromosome segregation and cell survival
- RNA processing machineries in germ cells
- the regulation of hematopoietic cell activation through novel signaling networks linked to the F-actin cytoskeleton
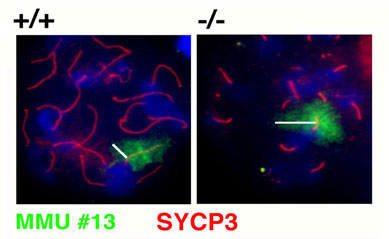
The first project aims at understanding the contribution of SMC (structural maintenance of chromosomes) proteins and their complexes to essential processes in mitosis and meiosis such as sister chromatid cohesion and segregation, DNA recombination and repair, chromosome structure and behaviour. We were first to isolate mammalian SMC proteins and to implicate them in repair of certain DNA damage. The role and mechanisms of SMC complexes and their regulatory factors in DNA repair is still incompletely understood. In our studies on germ cells, we identified a novel SMC protein that in a mouse model turned out to be essential for meiotic sister chromatid cohesion, telomere integrity, and chromosome structure. Our goal in this area is to further elucidate the function of SMC complexes in male and female meiosis, and their contributions to avoiding aneuploidies such as the frequent trisomie syndromes in humans.
In male germ cells we are also interested in RNA dynamics, which is undescribed in many respects. We currently focus on a tudor domain protein, which is required for spliceosome assembly and nonsense-mediated RMA decay in spermatocytes and spermatids.
In our hematopoietic project area we try to understand factors involved in the B cell switch to expression of specific immunoglobulin isotypes such as IgE, which is key to the allergic reaction. We further aim at deciphering the F-actin cytsokeletal dynamics of hematopoietic cells, important for migration, adhesion, homing.
Future Projects and Goals
In all of our research areas we use integrated approaches, which include a wide range of techniques like biochemical, cellular, genetic, and organismal methods, to deepen our understanding of central processes in mammalian biology, which are highly significant for human health. Recently, we added a focus on the role of chromosome-associated proteins in cancer. Thus, we will develop additional mouse models in both areas, will design and use further molecular assays, to address functional and mechanistic problems. The immediate focus is on SMC proteins, their high-molecular weight complexes and regulators in mitosis and meiosis, on TDRD6 in male germ cells, and on SWAP-70 and related proteins in the hematopietic cell area.
Methodological and Technical Expertise
- Mammalian spermatocyte and oocyte methods including oocyte injection, live imaging, and chromosome analysis
- Large variety of immunological methods addressing mammalian lymphoid and myeloid cells
- Analysis of F-actin cytoskeletal dynamics, integrin activity, cell migration and other cell biology parameters
- Protein biochemistry methods
- Analysis of DNA recombination, DNA repair, DNA binding proteins